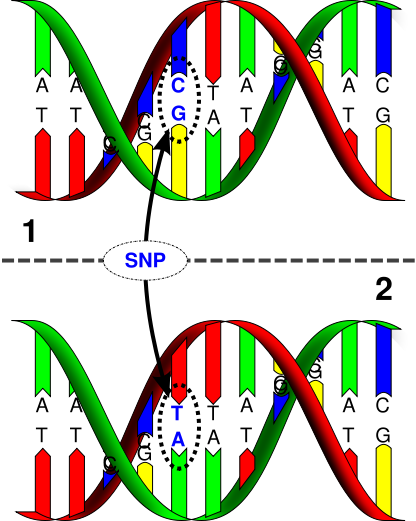
SNP (pronounced snip) stands for single nucleotide polymorphism. SNPs are locations within the human genome where the type of nucleotide present (A,T,G, or C) can differ between individuals. SNPs are the most common type of genetic variation found among people. At least 1% of a population must contain the same nucleotide variation for it to be considered a SNP. SNPs occur roughly every 300 nucleotides, and since there are 3 billion nucleotides in the human genome, there are approximately 10 million SNPs. Over 99% of the genome is identical between individuals, so SNPs provide researchers a way to study the genetic root of the differences that are apparent across the human race. Although most SNP’s have no effect on health, they can be important tools for genetic research. For example, SNPs are the basis for genome wide association studies that allow researchers to determine regions of the genome that may be important in disease development. The identification of disease causing mutations is an important first step in developing a therapy. Since SNPS are hereditary and shared by individuals of common descent, they can also be used as a way to track ancestry. To this end, the International HapMap Project was founded in 2002 with the goal of locating particular SNPs within the human genome that can serve as “tags†for studying human variation and disease. All of the data generated by the International HapMap Project is freely available.
CLICK HERE to learn more about DNA
